A New Protocol for Atomic-Level Protein Structure Modeling and Refinement Using Low-to-Medium Resolution Cryo-EM Density Maps
![PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/682fc6a2dcf6edaadacb1a2f90fdd137aef28d15/2-Figure1-1.png)
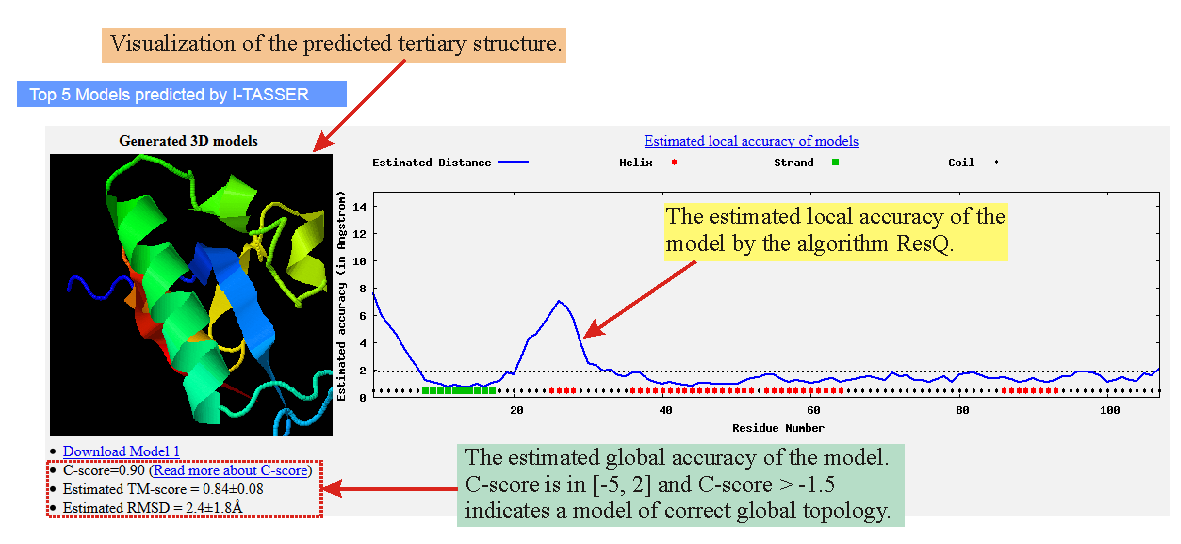
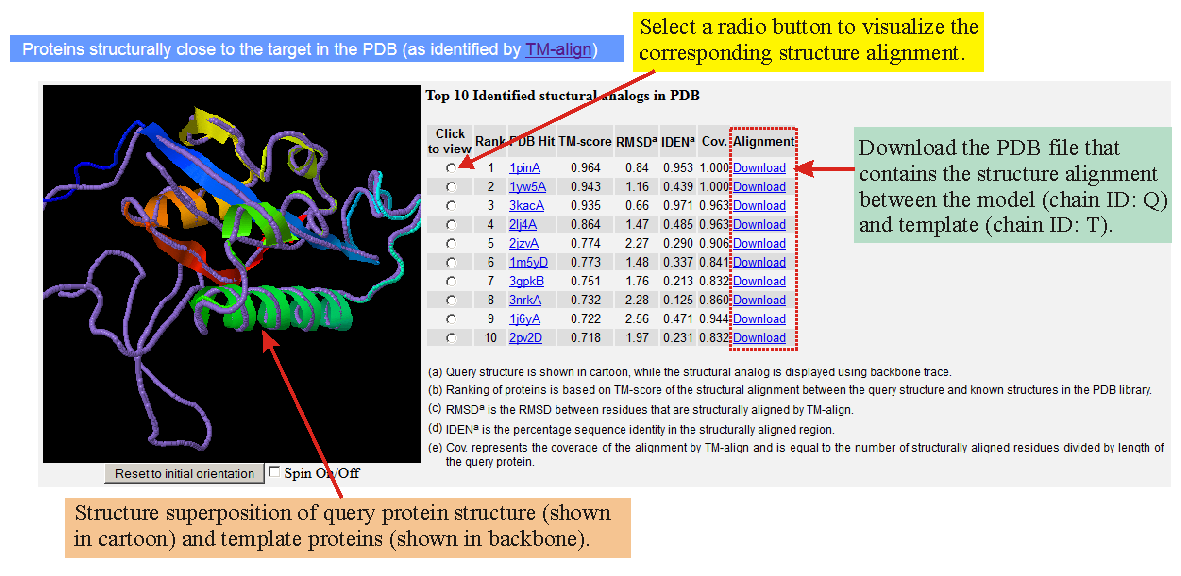
PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar

Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations. - Abstract - Europe PMC
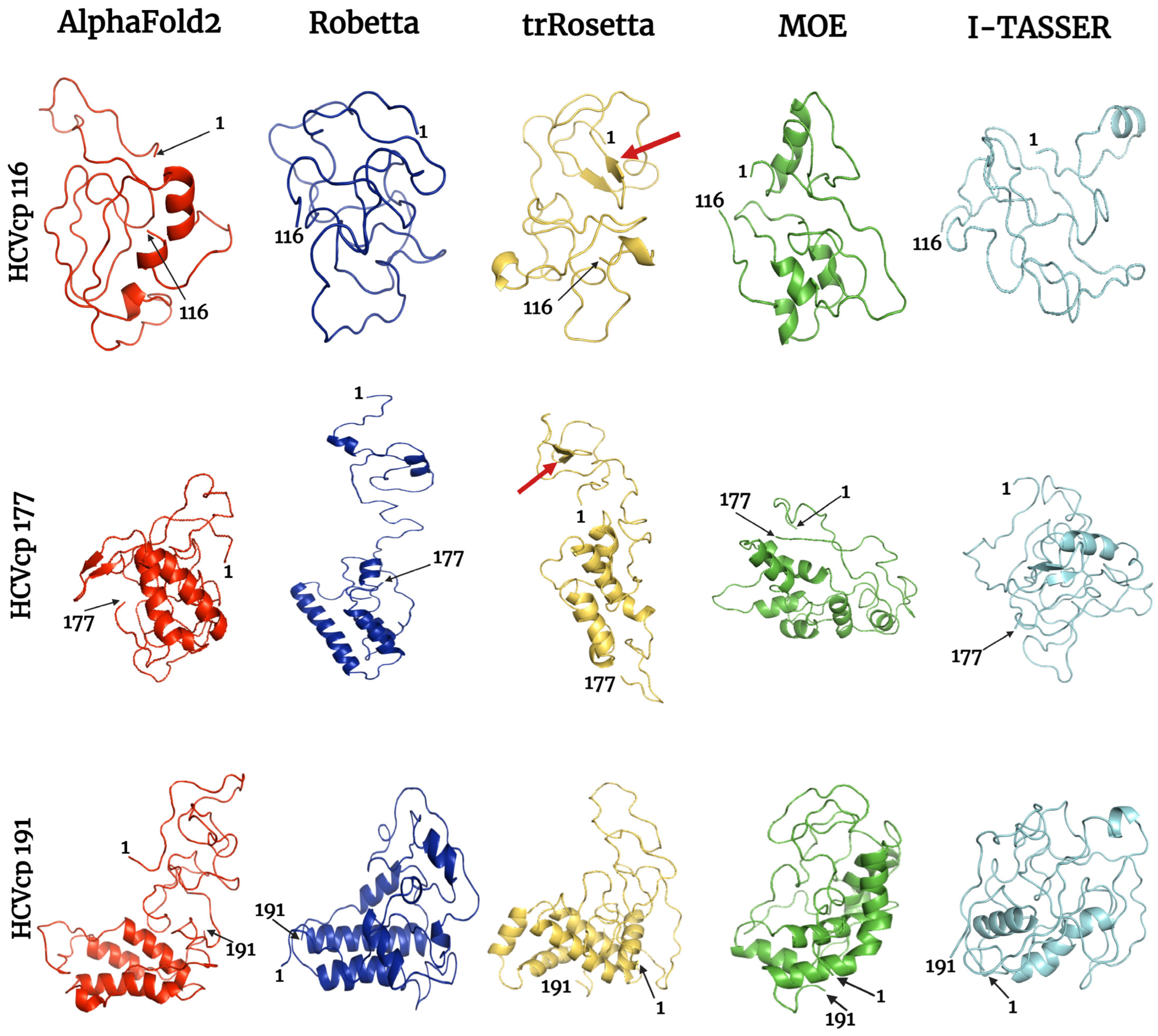
Bioengineering | Free Full-Text | Comparison, Analysis, and Molecular Dynamics Simulations of Structures of a Viral Protein Modeled Using Various Computational Tools

I-TASSER-MTD: a deep-learning-based platform for multi-domain protein structure and function prediction | Nature Protocols
Bioinformatics |3D Protein Structure Prediction |I-TASSER Result Analysis of Protein Structure - YouTube
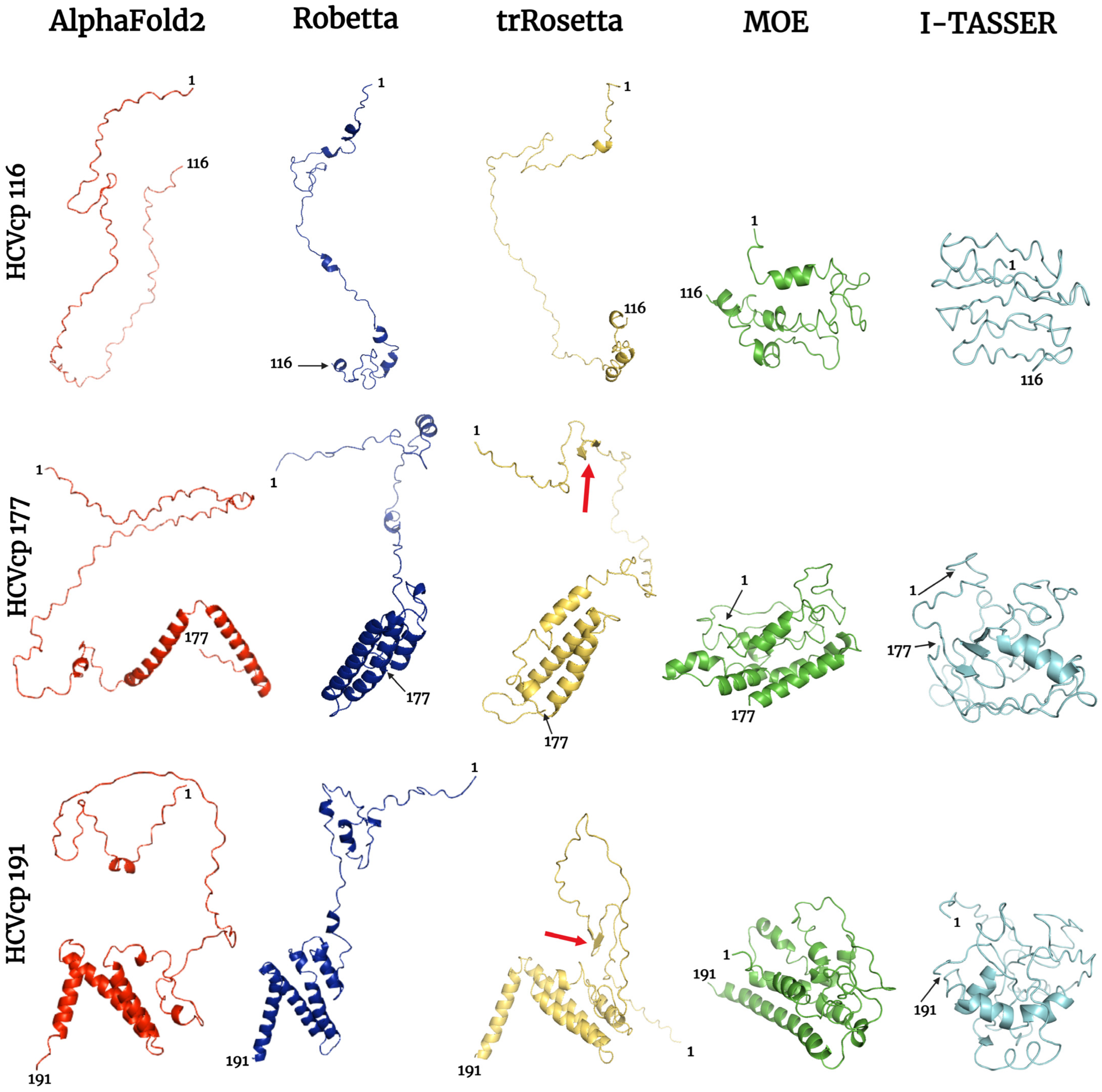
Bioengineering | Free Full-Text | Comparison, Analysis, and Molecular Dynamics Simulations of Structures of a Viral Protein Modeled Using Various Computational Tools

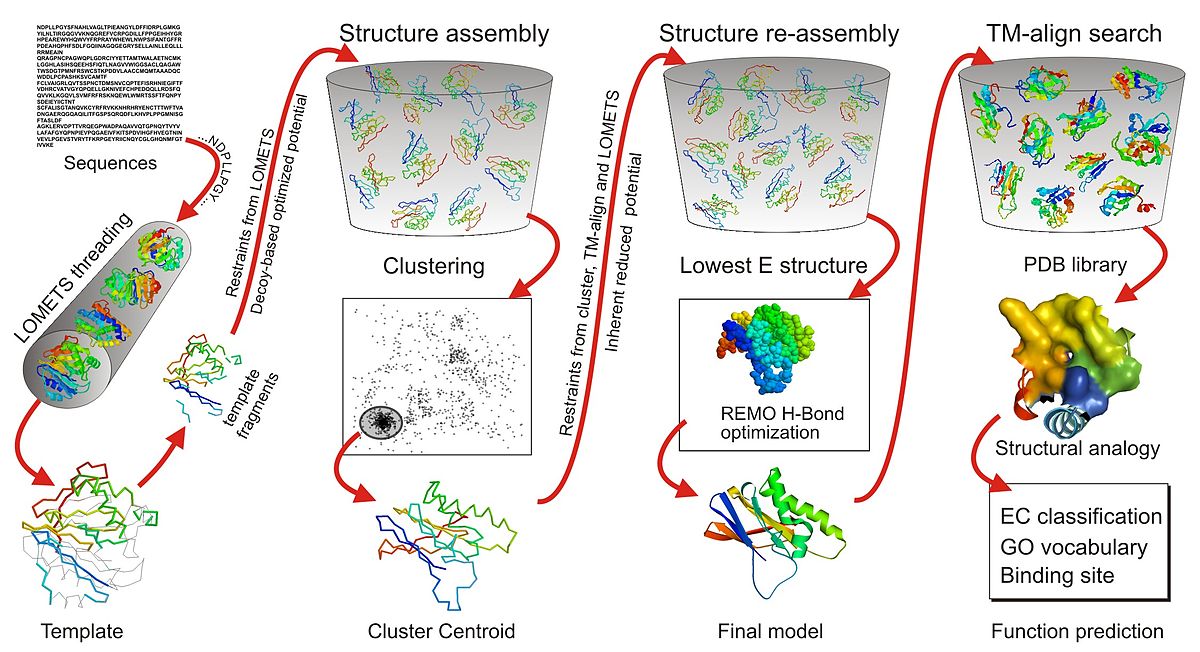

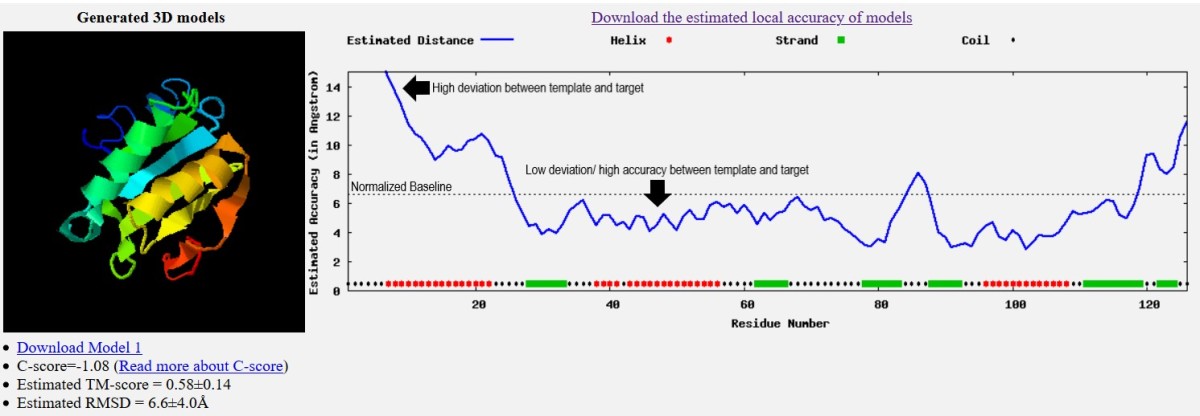

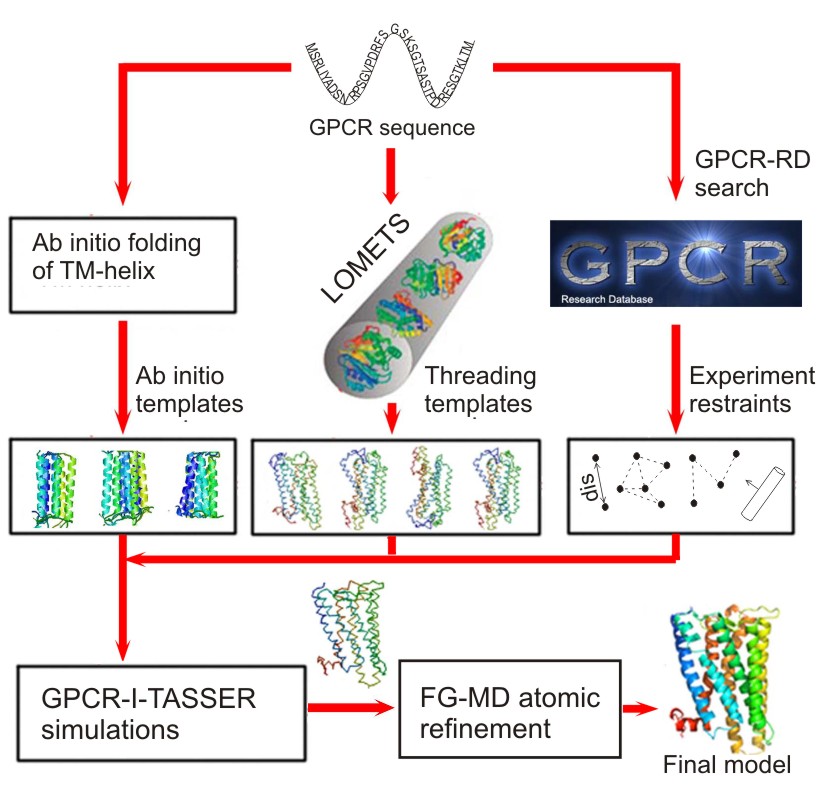
![General workflow of I-TASSER for protein structure prediction [30] | Download Scientific Diagram General workflow of I-TASSER for protein structure prediction [30] | Download Scientific Diagram](https://www.researchgate.net/publication/281540998/figure/fig1/AS:281366399864834@1444094385414/General-workflow-of-I-TASSER-for-protein-structure-prediction-30.png)